Last year over 700 boat owners who have non-trailered boats moored in marinas around Aotearoa…
Harnessing decay rates for coastal marine biosecurity applications: A review of environmental DNA and RNA fate
The recent study led by PhD student Michelle Scriver focuses on molecular techniques based on environmental DNA and RNA (eDNA/eRNA) in marine environments and their application in biosecurity programs. Environmental nucleic acids (eNAs) have gained popularity as a noninvasive tool for detecting non-indigenous species (NIS) and monitoring biodiversity. However, uncertainties regarding eNAs detection probabilities and the source population’s location hinder the widespread adoption of this tool in marine biosecurity.
To address these uncertainties, the study reviews 20 recent reports on the degradation mechanisms and fate of eNAs in the marine environment. The researchers classify the critical factors that influence eNAs’ persistence and outline the complex interaction between degradation processes and environmental conditions. The aim is to contribute to the development of knowledge in this area and provide guidelines for accurate decay rates, which can be used to build effective marine biosecurity tools for target detection and biodiversity assessment.
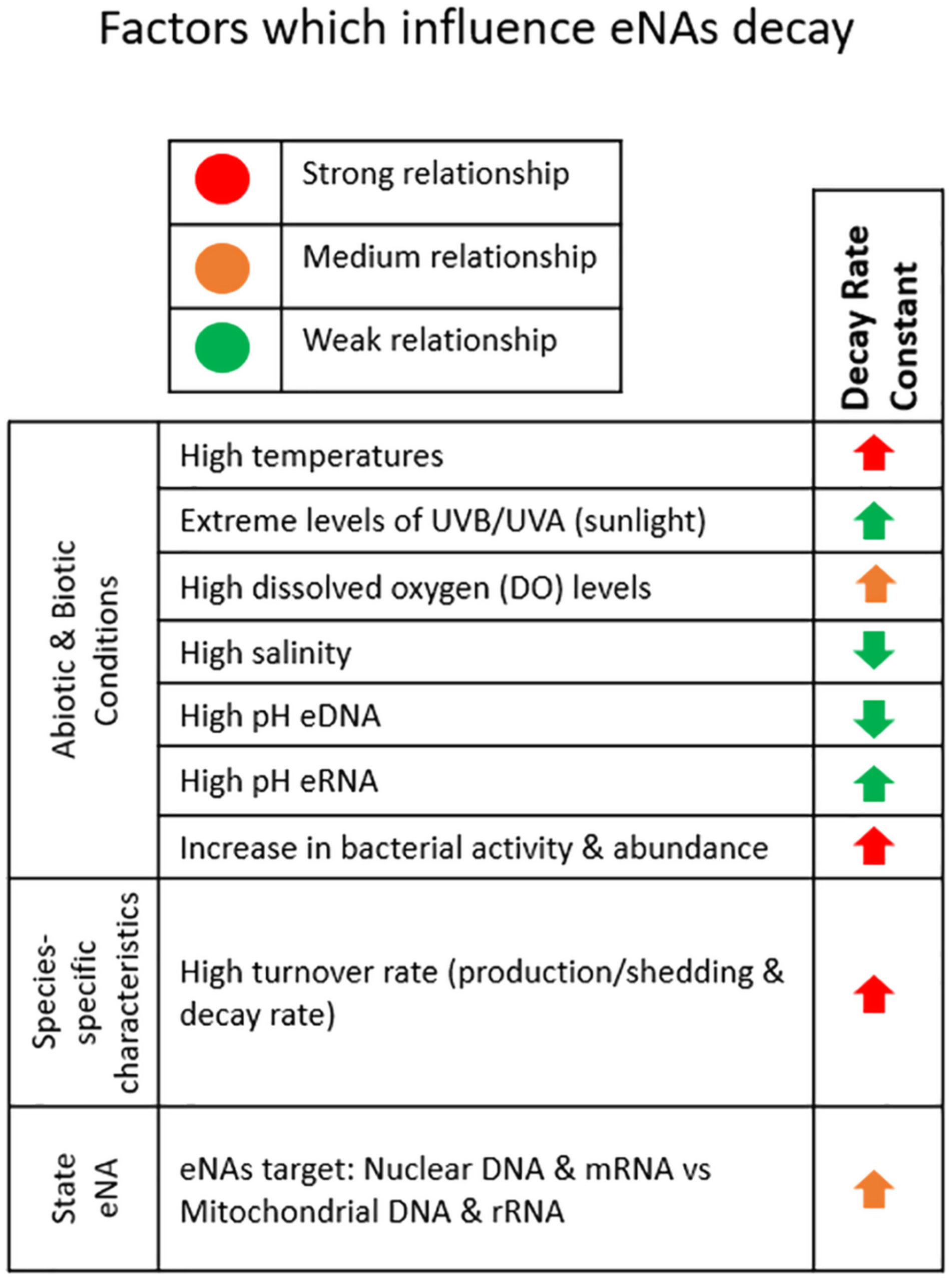
The manuscript emphasizes the importance of understanding the fate of eNAs, including changes in their state, persistence, and degradation. The decay rate or decay constant is a key parameter used to describe the rate at which eNAs concentration decreases over time. The study reviews factors that influence eNAs degradation in coastal marine habitats and summarizes marine decay rates of eDNA/eRNA from the literature.
The review focuses on coastal regions as they are biodiversity hotspots and prone to marine species incursions. The authors compile a comprehensive summary of eNAs decay rates organized by taxonomic class to provide researchers and biosecurity management programs with insights into the expected decay rates for targeted species. The study highlights the need for further research on how decay rates influence metabarcoding data and the ecological interpretation of such datasets.
While the manuscript provides recommendations for decay rates and their integration into biophysical models, it acknowledges the importance of considering other factors and characteristics of eNAs and oceanographic data. These factors include resuspension, transport, dilution, settling, diffusion, stratification, and advection. Developing frameworks and validating biophysical models in the field with established source populations are crucial for accurately predicting eNAs detectability and distribution.
Advancing these molecular tools within marine biosecurity programs requires collaboration among experts from various fields, including oceanographers, computer scientists, and scientists studying eNAs ecology. Continuous communication and informed decision-making will contribute to protecting marine resources and minimizing harm to native ecosystems.




Comments (0)